Reprogramming, stem cells and oncogenesis

Objectives
Projects
Axis 1: Deciphering the reprogramming roadmaps toward pluripotency and malignancy
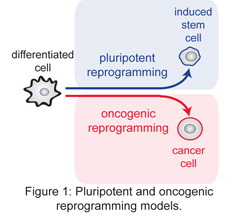
Dedifferentiation and cellular plasticity are key features of pluripotent and oncogenic reprogramming (Figure 1). Such plasticity is regained when the epigenome of a somatic cell is induced to dedifferentiate and reprogrammed toward the pluripotent state by the transcription factors Oct4, Sox2, Klf4 and c-Myc (iPS cells – induced pluripotent cells). The iPS cells generation process opened new avenues for regenerative medicine but the approaches are still limited by the poor efficiency of the procedure and by the wide differentation potential of the resulting iPS cells. Understanding the molecular mechanisms triggering the early steps of PR is crucial for the efficient generation of high-quality iPS cells.
Plasticity features are also re-acquired at multiple stages of oncogenesis, with the example of oncogenic reprogramming induced by mutated Ras and c-Myc that involves dedifferentiation and reacquisition of developmental programs. However, even if oncogenic reprogramming and dedifferentiation are considered as fundamental steps in cancer initiation, the molecular networks precluding the conversion of a somatic cell to a tumorigenic state remain unclear. The development of models recapitulating the early steps of oncogenic reprogramming is crucial to broaden our understanding of cancer initiation.
The main objective of the team is to identify novel factors/mechanisms governing dedifferentiation. We are particularly deciphering the initial steps triggering the reprogramming of a somatic cell toward the pluripotent and/or oncogenic states. To do so, we developed in the past years large-scale approaches and original genetic models to tackle such biological questions. Our efforts recently led to the identification of the Netrin-1 signalling pathway as a novel reprogramming roadblock (Lavial F. et al., patent 2014 and Ozmadenci D. et al., Nature Communications 2015) and the Bcl11a and Bcl11b transcription factors as regulators of cell identity (Huyghe A. et al., submitted).
Axis 2: Control of pluripotent stem cell self-renewal
-
Fabrice Lavial
Team leader